Contents
Setting up a statistical design in SPM2
When you are familiar with the SPM2 statistics interface, you might consider using SPM2 batch mode for statistics, as the model setup is GUI intensive and prone to error.
An example dataset
In the following example, we will set up the model for one session from the MarsBaR example dataset. This is a simple one-condition, one-session event-related design - see the MarsBaR tutorial, and the README file in the dataset archive for more detail. Here we will analyze session 3 of that dataset. To follow this example, you may want to download and unpack the data archive somewhere, and run the "run_preprocess" script in the "batch" directory in the archive; again, the MarsBaR tutorial lists these steps in more detail. Next you may want to make a new directory to store your example SPM analysis. Change to this directory in matlab ("cd my_directory" or click on the SPM "Utils" menu, and choose "CD" to browse to a directory with the GUI).You should also have a look at the various example datasets available on the SPM FIL pages.
Specifying the design
To run the GUI, click on the fMRI button in the SPM2 buttons window.
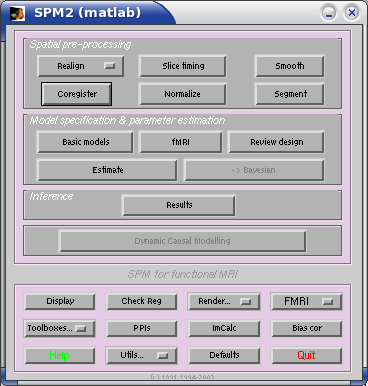
At the prompt "specify design or data", select "design". Give 2.03 as the interscan interval, and 126 as the scans per session. Note that, if we had been analysing - say - three sessions, the entry might have been "126 126 126". Next choose to specify the design in scans rather than seconds. It just so happens that the onset file that I have is in TRs, but of course your onsets could be in seconds from the beginning of the scan run, in which case you would have chosen "seconds" here.
Basis set
Choose: hrf (with time derivative). The basis set is the set of functions that can be used to model your events or blocks. More complex basis sets use more degrees of freedom, can be confusing to review, and may well be better at capturing differences in shape of response between events, or from an assumed HRF shape. Note that, unlike SPM99, you are forced to choose the same basis set for all the events in your model. This design involves a simple visual response, which I expect to be more-or-less the same shape as the standard HRF, so I suggest you choose "hrf (with time derivative)" as a basis set here.
Model interactions (Volterror)
No, no, a thousand times no.
Number of conditions/trials
Choose 1 - we only have one event-type.
Name for condition/trial 1?
Enter: Flashing checkerboard
Vector of onsets
for "Flashing checkerboard". We need to enter the onset times, in TRs (because we chose TR units above), of the flashing checkerboard events. The beginning of the first TR is 0. Here are the event onsets as they should be typed into the entry box:
1.0 9.6 10.2 13.7 22.0 24.0 25.2 30.7 38.2 46.3 47.8 51.6 52.8 58.6 59.9 63.9 79.6
You can copy and paste these out of the browser window, to save time.
Duration(s) (events=0)
The default event duration for SPM2 is "0" - which will actually be converted to an assumed event duration of about 1 second. If each event has a different duration, you will have to enter one value here for each onset, but our events are all the same duration, and the default duration is OK, so enter a single 0.SPM2 does not distinguish events and blocks in its modelling; a block is simply an event with a long duration.
Parametric modulation
Choose: None. This option allows you to model the fact that different events may have different weights; for example you could imagine an experiment where the flashing checkerboard had been of different intensities for different events. This option allows you to enter a weight for each event to take this into into account, but that is not the case here.
User specified [regressors]
Choose: 0. Here you can enter any more regressors for this session. The most common use of this option is to enter the movement parameters into the model, to try and soak up residual movement-related variance. To do this, you could click on "1" (= 1 block of regressors) and then, when asked for values for regressor 1, type "spm_load" in the input box. This allows you to select the rp_*.txt file for this session, and return the values in the file as the regressors. SPM works out that there are in fact 6 regressors, and asks you to name these (say "translation x", "translation y" etc). For simplicity we are not doing that for this model.After all this is done, you will have a new "SPM.mat" file in your matlab current directory, containing the design.
Adding data to the design
Now you have configured the design, you need to specify the scans to use with the design. Click on the "fMRI" button again. This time, select "data" to specify. Next select the "SPM.mat" file you have just created. Next, select the 126 scans for the design; navigate to the "sess3" directory in the example dataset, and select the "snura_sess3_0*img" files. (If you can't see these files, maybe you have not run the "run_preprocess" batch script from the MarsBaR dataset?)
Remove Global effects
Choose: none. This give you the option of using proportional scaling on your data. This will have the effect of dividing all the voxel values for each scan by the thresholded mean voxel value for that scan, so the voxel values are now a ratio of their mean; see the SPM statistics tutorial for more details. This option can be useful in scanners with significant fluctuations in signal over the course of the scanning session - and specifically, fluctuations that are of higher frequency than your high-pass filter (see below). The disadvantage is this can lead to odd effects if the global signal in the scans has been affected by the activation signal. See Kalina Christoff's global scaling page for more details. The Bruker scanner seems to have fairly good signal stability, and we generally do not use proportional scaling for our data. Some people have found that global scaling is more useful for Oxford data.
High-pass filter
Choose: specify; 128. Here we enter the wavelength in seconds for the high-pass filter. The high-pass filter passes high frequencies, but eliminates low frequencies. FMRI data has a very disproportionate amount of noise at low frequency, and most FMRI designs have small amounts of low frequency components in them, so it is efficient to remove the low frequencies entirely with a high-pass filter. In fact, the SPM2 autocorrelation options more-or-less require that you have a high-pass filter, because they can only model a very limited range of autocorrelation, and this range is suitable for FMRI time-courses after high-pass filtering. The choice of high-pass filter is rather difficult. One way is to look at the design and the data frequency power spectra to see if there is a cut-off that eliminates much of the noise while leaving almost all of the model frequencies. MarsBaR has a tool to do this in an ROI, for example. Here I suggest you accept the default, which captures much of the worst noise. Note that SPM2 does not attempt to match the suggested high-pass cutoff to your design, as was the case for SPM99.
Correct for serial correlations
Choose: AR(1). If you select this option, SPM will attempt to estimate the autocorrelation in the data, and adjust for it - by so-called "whitening" of the time course. This step is necessary if we want to be able to interpret our fixed-effect (e.g single-subject) p values. Because FMRI data with a short TR (as here) have some autocorrelation, we have to take this into account in calculating the statistics. SPM does this using a two-pass procedure. First it estimates the model without taking account the autocorrelation. In the process, it selects out all voxels having a large amount of variance explained by the design (in fact, by the effects of interest). It then pools the covariance of all these "activated" voxels, and estimates a best-fit model of their autocovariance, using a very constrained model. By default this allows the data to have an autocorrelation from an AR process having more or less an AR(1) coefficient of 0.2. If your actual AR(1) coefficient if far off this, you are out of luck, and will have to try and get SPM to use a model that is closer to your data. There are other problems: by pooling the autocorrelation, SPM assumes the form of the autocorrelation is the same across all "activated" voxels, which may well not be the case. For example, if you have included regressors for movement, voxels with high movement-related variance will be included as activated, and may change the autocorrelation values. After SPM has calculated the autocorrelation, it calculates a filter that will remove the (calculated) autocorrelation (this is the "whitening" process), and then reruns the model using this new filter.
Estimate the design
Now, you have configured your design. To estimate the design, click on the Estimate button, and select the "SPM.mat" file in your current directory. You're done. Now all you need to do is interpret the data. What could be easier than that?
